Identification & Classification of Aminoacyl tRNA Synthetases
Aminoacyl-tRNA synthetases (aaRSs) play a key role in the process of protein biosynthesis in all organisms. They are responsible for the fidelity of translation process by the precise recognition and correct aminoacylation of cognate tRNA molecules. There are twenty aminoacyl tRNA synthetases found in all organisms and each one is specific for single amino acid. [See link]
Aminoacyl-tRNA synthetases are organized into two classes, based on their active site architectures -
Class-1: MetRS, ValRS, LeuRS, IleRS, CysRS, ArgRS, GluRS, GlnRS, TyrRS, TrpRS
Class-2: SerRS, ThrRS, AlaRS, GlyRS, ProRS, HisRS, AspRS, AsnRS, LysRS, PheRS
Aminoacyl tRNA synthetases differ in amino acid sequence length, 3D structure, molecular weight, subunit organization and have limited sequence homology. Prediction of aaRSs is very important for the functional annotaion of proteomes. We have developed "icaars" prediction method of aaRSs by using hybrid approach of Support Vector Machine (SVM) and PROSITE.
1. Prediction of aaRSs: First we have developed prediction method between aaRSs and non-aaRSs and achieved 92.83% accuracy with 0.86 MCC for 40% non-redundant datasets by using hybrid approach2 [SVM & PROSITE].
2. Prediction between class-1 & class-2 aaRSs: We further classified the Aminoacyl-tRNA synthestases into Class-1 and Class-2 aaRSs and achieved 97.47% acuuracy with 0.95 MCC for 40% non-redundant datasets by using hybrid approach2 [SVM & PROSITE].
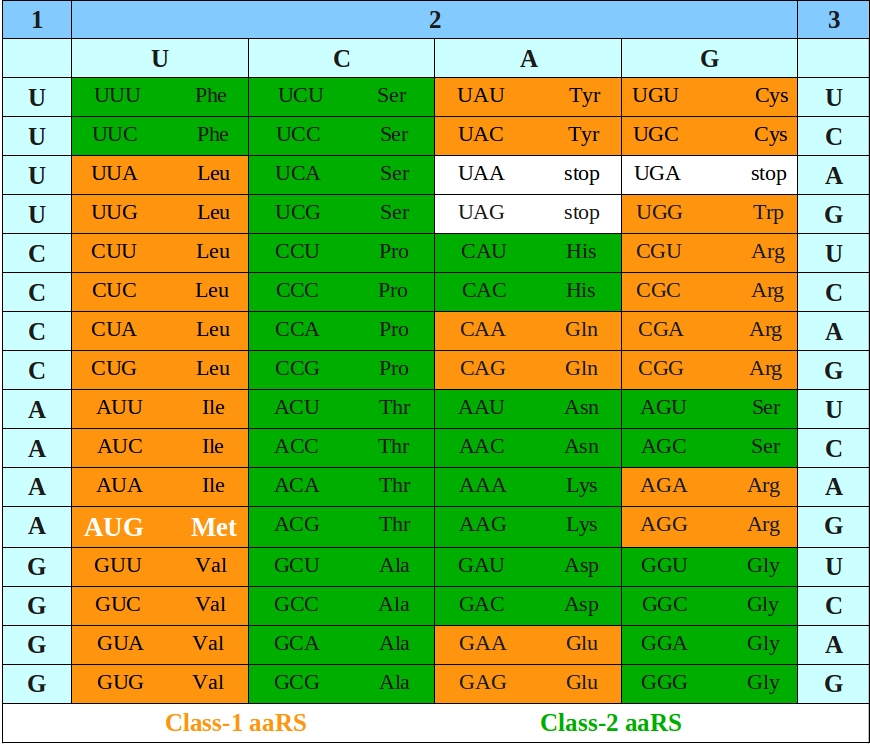
Figure 1 - Class-specific representarion of aaRSs in Codon-table

Figure 2 - Hann diagram of amino acids for class-specific aaRSs
If you are using this webserver, please cite:
Panwar, B and Raghava, GPS (2010) Prediction and classification of aminoacyl tRNA synthetases using PROSITE domains. BMC Genomics 2010, 11:507.