Publication reference::This field have the published reference of peptide in the coded form.The first of code specify the first author sirname. The next two digits specify the year of publication.The next word after underscore denotes the journal name.The last digits after second underscore denotes stating page of published reference.
The deciphacialed publiaction reference:
first author of paper along with last two digits of year of publication_journal code_first page of published reference.
This is field is internally linked to flat files which is further hyperlinked to NCBI.
Schematic diagram of B-cell epitope
Statistics on "pathogen group' and 'immunogenicity' vice distribution of B cell epitopes in the database
| Pathogen |
Immunodominant |
Immunogenic |
Non-immunogenic |
| Virus (2046) |
415 |
1474 |
157 |
| Bacteria (539) |
130 |
159 |
250 |
| Protozoa (236) |
139 |
57 |
40 |
| Fungi (53) |
17 |
23 |
15 |
| Others (157) |
62 |
84 |
9 |
Application of Bcipep data in prediction methods
The Physio-chemical properties of amino acids in B-cell epitope can be identified e.g., Hydrophilicity, flexibility, accessibility, turns and tested on this experimental determined epitopes.
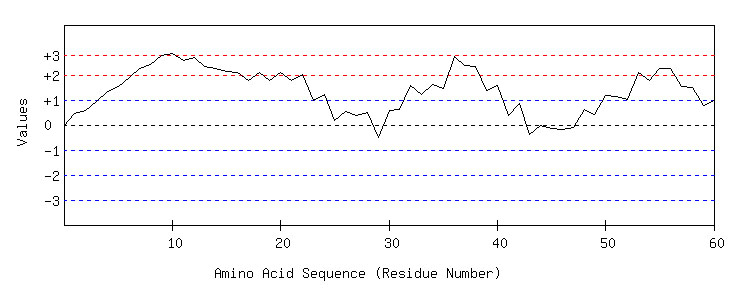
Seq =AAAHHG
STEERENANPNANPNANPGFGFERRWEQ
SADADHGFGFDWERWNANPNANPDSF
Diagram showing Hydrophilicity scale on Y axis and amino acids in the x axis.
The amino acids in the protein above the threshold value (>2) of a physio-chemical scale (eg., Hydrophilicity ) can be predicted as B-cell epitope. Bcipep can be helpful in determining threshold value, analysis and validation of the existing methods for the prediction of B-cell epitope. New method(s) can be used like machine learning language which can differentiate between epitope and non-epitope.
B-cell epitopes obtained from this database (Bcipep) have been used for evaluation of prediction methods of continuous B-cell epitopes in antigenic sequences using physico-chemicalproperties.
see the website: BcePred: webs.iiitd.edu.in/raghava/bcepred/
Reference:
Saha.S and G.P.S.Raghava. BcePred:Prediction of Continuous B-cell Epitopes In Antigenic Sequences using physico-chemical Properties. In G.Nicosia,V.Cutello,P.J.Bentley and J.Timmis(Eds.) ICARIS 2004, LNCS 3239, pp. 197-204, Springer,2004.
Bcipep data mangement
All data in Bcipep has been maintained in "Relational Database
Management System" (RDBMS) called PostgreSQL which is a public domain software freely available to public. Full information
about a peptide has been stored in a single table. Related information
like publication reference and source proteins reference has been stored in directories having internal links to main database.
Specification for Access of Bcipep
The Bcipep web server was developed in a UNIX environment on SUN server 420E in Solaris 7.0. This server is designed to provide easy
access to the user, based upon a set of simple GUI (Graphical User Interface)
forms. Methods for searching the databases and displaying the selected
objects were built with a combination of HTML, CGI-scripts in PERL
5.4.
Requirements for accessing Bcipep on your system
-
Windows 95 and above for personel computers
-
Internet Explorer 4.01 and above/Netscape 3.01 and above
Best view of Bcipep can be observer with
-
a 1024x768 screen
-
not to override the web pages with your windows default colors.
-
to set font size of browser at 12 (Helvetica) or medium.