Citation: Thakur N, Qureshi A, Kumar M (2012)
AVPpred: collection and prediction of highly effective antiviral peptides. Nucleic Acids Res. 2012 Jul;40(Web Server issue):W199-204. doi: 10.1093/nar/gks450.
PMID:
22638580
Antiviral Peptide Prediction Server
 |
-
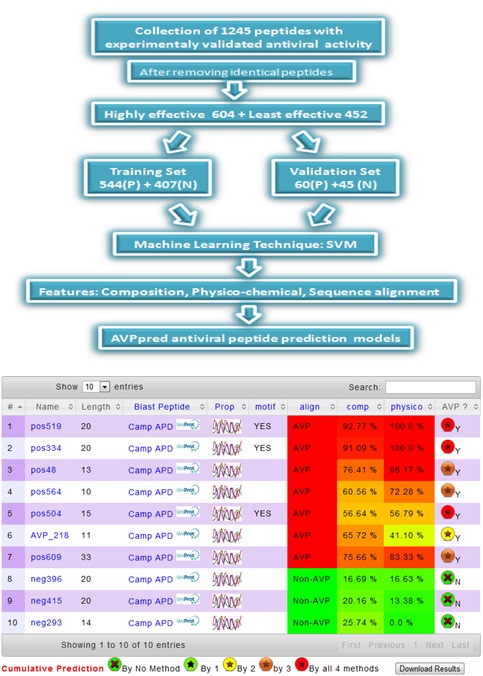
SVM based models using peptides sequence features; Antiviral peptide Motifs, sequence alignment, amino acid compositions and physiochemical properties.
-
Highly user friendly web interface
-
An option to generate overlapping peptides of desired length from protein sequence
-
AVP predictive scores output for all the models is provided against each peptide in numerical as well as graphical form using different coloring scheme to help the user to interpret the results to select the highly effective antiviral peptides.
-
An analysis tool blast to search the similar peptides in the CAMP, AMP databases and Uniprot.
-
AVPpred would certainly be helpful to researchers working on peptide based antiviral therapeutics development
Related web servers:
- AVP-IC50Pred
 -Prediction of peptide antiviral activity in terms of IC50. -Prediction of peptide antiviral activity in terms of IC50.
- HIPdb
 -Database of HIV inhibiting peptides. -Database of HIV inhibiting peptides.
- AVPdb
 -Database of antiviral peptides. -Database of antiviral peptides.
|
Bioinformatics center IMTECH chandigarh

