Algorithm
Peptide Properties
To discriminate the peptides having Quorum sensing activity from non-Quorum sensing we have used following features.
1. Amino acid composition+ Dipeptide composition
2. Binary pattern for N- and C- terminal
3. Physico-chemical parameters including secondary structure, charge, size, hydrophobicity and amphiphilic character as these yielded an appreciable accuracy using machine-learning technique. The values of physico-chemical properties were retrieved from AA index database (Kawashima and Kanehisa 2000)
4. Amino acid composition+ Dipeptide composition+ Binary pattern for N- and C- terminal+ Physico-chemical parameters
Machine Learning Techniques
Support Vector Machine (SVM) was implemented using freely downloadable software package SVMlight (Joachims 1999, http://svmlight.joachims.org/). SVMlight is an implementation of Vapnik's Support Vector Machine (Vapnik, 1995) for the problem of pattern recognition.The software enables the user to define a number of parameters as well as to select from a choice of inbuilt kernel functions, including a radial basis function (RBF) and a polynomial kernel.
In addition we also used other machine learning techniques like Instance Based Classifier (IBk), Random forest from Weka package. Although the results from SVM were best. Random forests are an ensemble learning method for classification (and regression) that operate by constructing a multitude of decision trees at training time and outputting the class that is the mode of the classes output by individual trees. IBk is a K-nearest neighbors classifier. It can select appropriate value of K based on cross-validation and distance weighting.
Evaluation parameters
The performance modules constructed in this study were evaluated using a 10-fold cross-validation technique. In the 10-fold cross-validation, the relevant dataset was partitioned randomly into five equally sized sets. The training and testing was carried out ten times, each time using one distinct set for testing and the remaining nine sets for training. The performance of the methods was computed using the following formulas
Sensitivity (Sn) = [TP / (TP+FN)]*100
Specificity (Sp) = [TN / (TN+FP)]*100
Accuracy (Ac ) = [TP+TN / (TP+FP+TN+FN)]*100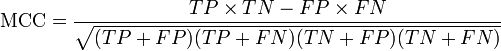
TP and TN are correctly predicted Quorum sensing and non-Quorum sensing peptides respectively.
FP and FN are wrongly predicted Quorum sensing and non-Quorum sensing peptides respectively.
ROC Plot
In order to evaluate performance of models using threshold-independent parameters, ROC (Receiver Operating Characteristic) were created for all models. ROC plots with area under curve (AUC) were created using ROCR statistical package available in R.
Motif Scan
In order to search QS motifs in user defined sequences motif scanning was done using MEME/MAST software (Bailey, T. L., M. Boden, et al.,2009).
Bailey, T. L., M. Boden, et al. (2009). "MEME SUITE: tools for motif discovery and searching." Nucleic Acids Res 37(Web Server issue): W202-208. Sing T, Sander O, Beerenwinkel N and Lengauer T (2005). “ROCR:
visualizing classifier performance in R.” Bioinformatics, 21(20),pp. 7881.
Kawashima, S. and M. Kanehisa (2000). "AAindex: amino acid index database." Nucleic Acids Res 28(1): 374.
Vladimir N. Vapnik, The Nature of Statistical Learning Theory. Springer, 1995.
Thorsten Joachims, Transductive Inference for Text Classification using Support Vector Machines. International Conference on Machine Learning (ICML), 1999.
SVMlight for Linux downloaded from http://download.joachims.org/svm_light/current/svm_light.tar.gz
Bioinformatics centre, CSIR-IMTECH chandigarh