Help
QSPpred is the first Web server for prediction and designing of Quorum sensing peptides. QSPpred comprises of various tools that will be beneficial for researchers in predicting Quorum sensing peptides and designing them. This page will help the users to explore QSPpred and speedup their research in field of Quorum sensing.
Result Interpretation
Three predictors QSPepPred, QSPepDesign, QSPepMap provide a user friendly and interactive output for easy interpetation and analysis.
Result will be analysed based on the color scale represented below. Color scheme shows least to most effective QS predicted peptides based on the MLTs (SVM,RF,KNN) scores, higher the score more the prediction confidence.
User can also download the result in Excel format.
QSPepPred: This tool will predict the input peptide sequence (fasta or multiple fasta) as Quorum sensing or non- Quorum sensing along with its corresponding Support Vector Machine (SVM) score and physico-chemical properties. Please provide input peptide sequences in the range of 5-50 aa in length. Peptide which is below and above the length range will be excluded.
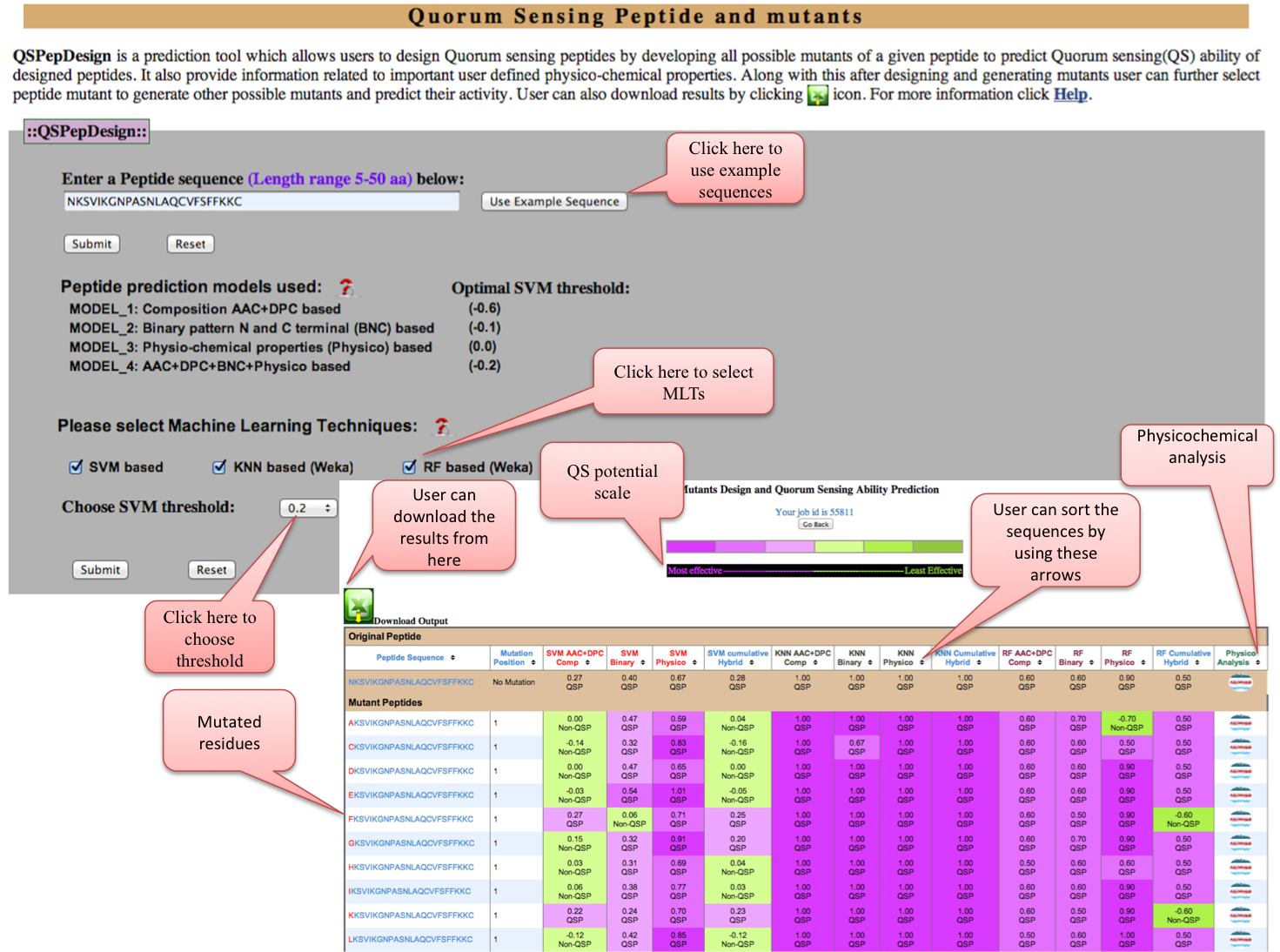
QSPepDesign: Designing of Quorum sensing peptides by developing all possible mutants of a given peptide along with predicting Quorum sensing ability of designed peptides can be done. It also provide information related to important user defined physico-chemical properties. Along with this after designing and generating mutants user can further select peptide mutant to generate other possible mutants and predict their activity. Please provide input peptide sequence in the range of 5-50 aa in length. Peptide which is below and above the length range will be excluded. Result will be analysed based on the SVM score, higher the SVM score more the prediction confidence.
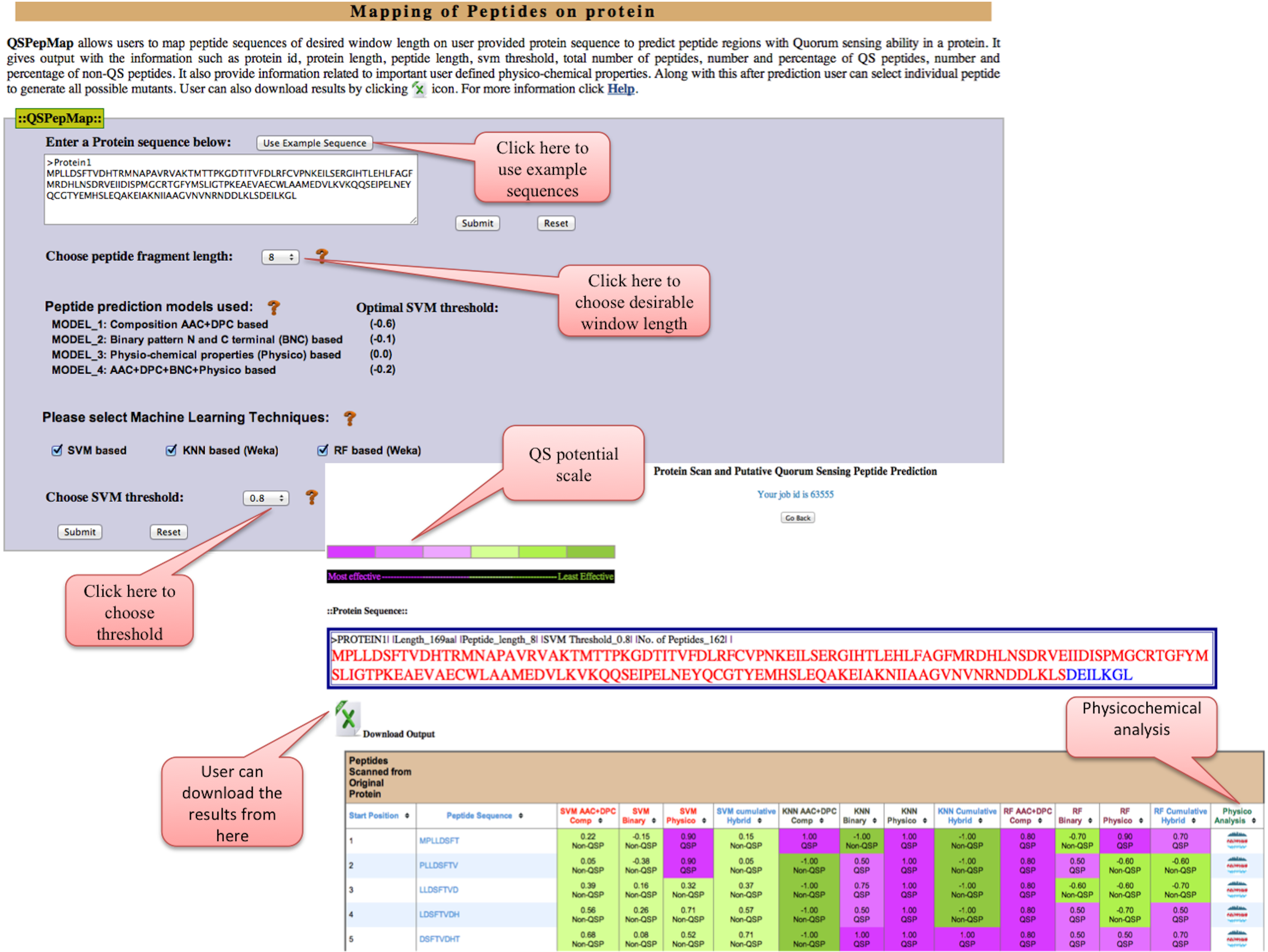
QSPepMap: This tool allows user to map QS peptide in protein sequence to predict the regions with Quorum sensing ability in a protein. User can get output information as protein id, protein length, peptide length, threshold, total peptides, number and percentage of quorum sensing and non-quorum sensing peptides. Information of some physico-chemical properties of individual peptides can also be explored.
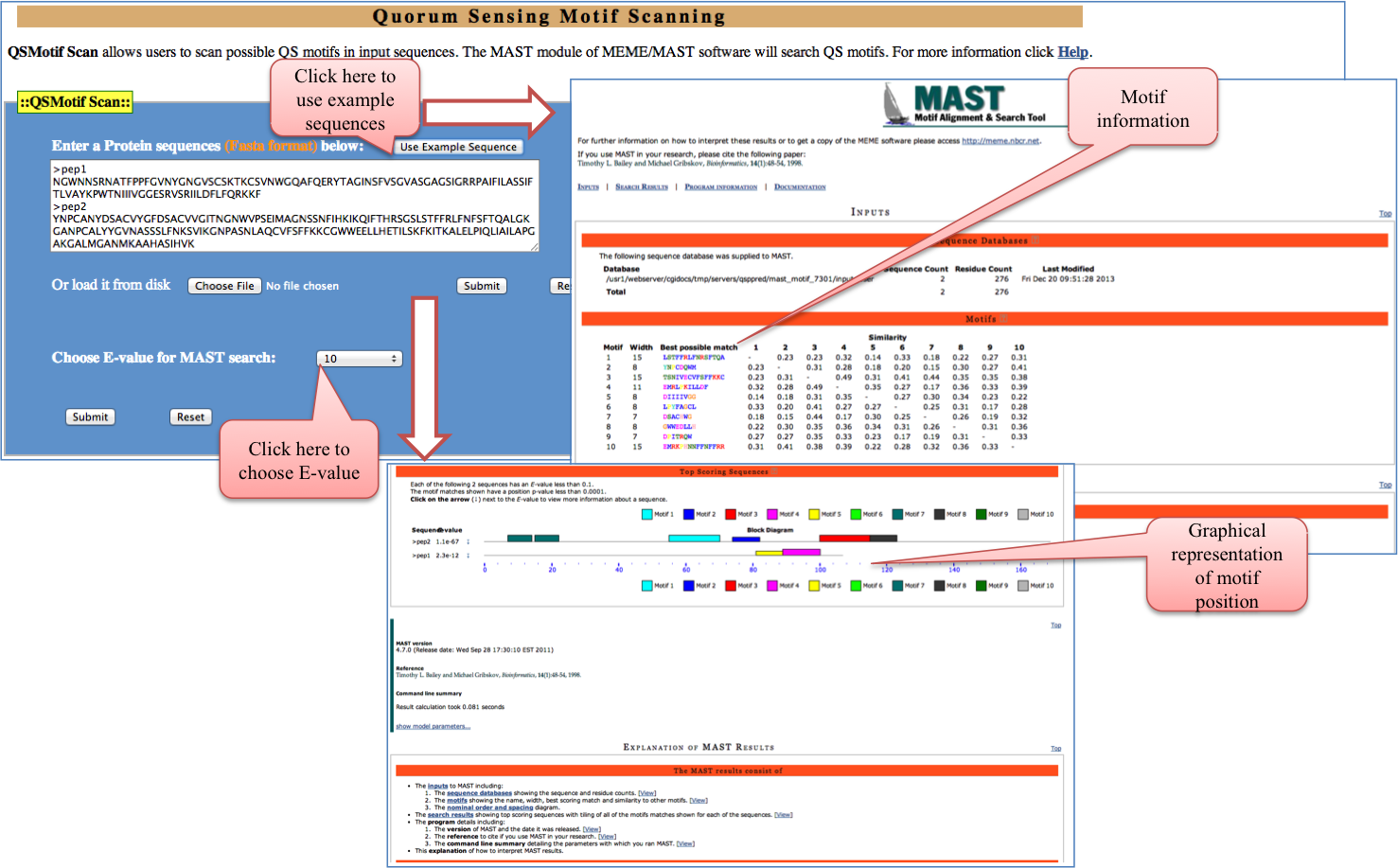
QSMotif Scan: In order to scan QS motif in input sequence at desirable E-value can be done using this tool. It scan motif by MAST module of MEME/MAST software.
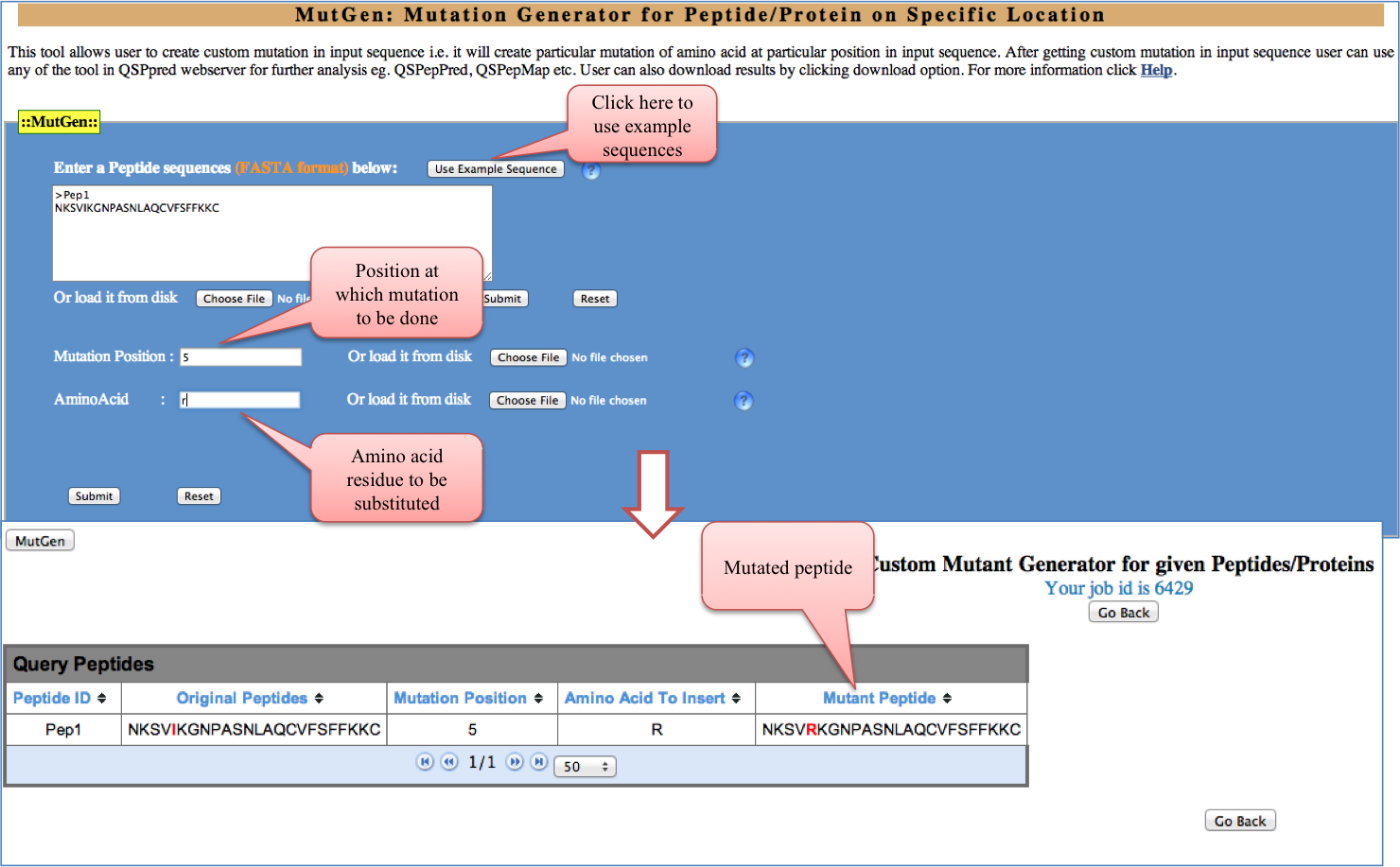
MutGen: If the user wants to generate custom mutation i.e. mutation at specific position and with particular amino acid in input sequence it can be done by use of this tool. PhysicoProp: This tool provides graphical representation of some important physico-chemical properties like percentage composition, hydrophobicity, preference for beta strand etc.
ProtFrag: If the user wants fragment of input sequence of desirable length and desirable overlapping length it can be done using this tool. Output can be further used as input for QSPepPred,QSPepDesign etc.
__________ __________ __________ __________ __________ __________ Most effective ------------------------------------------------------------ Least Effective
QSPepPred

QSPepDesign
QSPepMap
QSMotif Scan
MutGen
PhysicoProp

ProtFrag
